Python Read Last Line of Csv File
Most of the data is available in a tabular format of CSV files. It is very popular. You can convert them to a pandas DataFrame using the read_csv function. The pandas.read_csv is used to load a CSV file as a pandas dataframe.
In this article, y'all will learn the different features of the read_csv office of pandas apart from loading the CSV file and the parameters which tin can exist customized to become improve output from the read_csv function.
pandas.read_csv
- Syntax: pandas.read_csv( filepath_or_buffer, sep, header, index_col, usecols, prefix, dtype, converters, skiprows, skiprows, nrows, na_values, parse_dates)Purpose: Read a comma-separated values (csv) file into DataFrame. Also supports optionally iterating or breaking the file into chunks.
- Parameters:
- filepath_or_buffer : str, path object or file-like object Any valid cord path is acceptable. The string could be a URL too. Path object refers to os.PathLike. File-like objects with a read() method, such every bit a filehandle (e.g. via built-in open function) or StringIO.
- sep : str, (Default ',') Separating boundary which distinguishes between any two subsequent information items.
- header : int, list of int, (Default 'infer') Row number(s) to use as the column names, and the showtime of the information. The default behavior is to infer the column names: if no names are passed the beliefs is identical to header=0 and cavalcade names are inferred from the first line of the file.
- names : assortment-like List of column names to utilize. If the file contains a header row, then you should explicitly pass header=0 to override the column names. Duplicates in this listing are non immune.
- index_col : int, str, sequence of int/str, or Imitation, (Default None) Cavalcade(s) to use as the row labels of the DataFrame, either given equally cord proper noun or column index. If a sequence of int/str is given, a MultiIndex is used.
- usecols : listing-similar or callable Return a subset of the columns. If callable, the callable office will be evaluated against the column names, returning names where the callable function evaluates to True.
- prefix : str Prefix to add to column numbers when no header, e.1000. 'X' for X0, X1
- dtype : Type name or dict of column -> type Data type for information or columns. Eastward.grand. {'a': np.float64, 'b': np.int32, 'c': 'Int64'} Apply str or object together with suitable na_values settings to preserve and not translate dtype.
- converters : dict Dict of functions for converting values in certain columns. Keys can either exist integers or column labels.
- skiprows : list-similar, int or callable Line numbers to skip (0-indexed) or the number of lines to skip (int) at the start of the file. If callable, the callable function will be evaluated against the row indices, returning True if the row should be skipped and False otherwise.
- skipfooter : int Number of lines at bottom of the file to skip
- nrows : int Number of rows of file to read. Useful for reading pieces of large files.
- na_values : scalar, str, list-similar, or dict Additional strings to recognize as NA/NaN. If dict passed, specific per-column NA values. By default the following values are interpreted as NaN: '', '#North/A', '#N/A N/A', '#NA', '-1.#IND', '-ane.#QNAN', '-NaN', '-nan', 'i.#IND', '1.#QNAN', '', 'N/A', 'NA', 'NULL', 'NaN', 'n/a', 'nan', 'null'.
- parse_dates : bool or listing of int or names or listing of lists or dict, (default Simulated) If fix to True, volition attempt to parse the alphabetize, else parse the columns passed
- Returns: DataFrame or TextParser, A comma-separated values (CSV) file is returned as a two-dimensional data structure with labeled axes. _For total listing of parameters, refer to the offical documentation
Reading CSV file
The pandas read_csv office tin can exist used in dissimilar ways every bit per necessity similar using custom separators, reading just selective columns/rows and and then on. All cases are covered below one after another.
Default Separator
To read a CSV file, phone call the pandas role read_csv() and pass the file path equally input.
Step ane: Import Pandas
import pandas as pd Pace 2: Read the CSV
# Read the csv file df = pd.read_csv("data1.csv") # First 5 rows df.head() 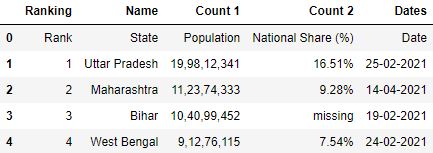
Dissimilar, Custom Separators
By default, a CSV is seperated past comma. Simply yous tin can use other seperators as well. The pandas.read_csvoffice is not limited to reading the CSV file with default separator (i.east. comma). It tin can exist used for other separators such equally ;, | or :. To load CSV files with such separators, the sep parameter is used to laissez passer the separator used in the CSV file.
Let's load a file with | separator
# Read the csv file sep='|' df = pd.read_csv("data2.csv", sep='|') df 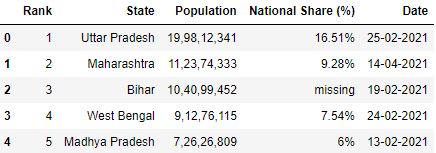
Gear up whatsoever row as column header
Let'southward come across the information frame created using the read_csv pandas part without whatever header parameter:
# Read the csv file df = pd.read_csv("data1.csv") df.head() 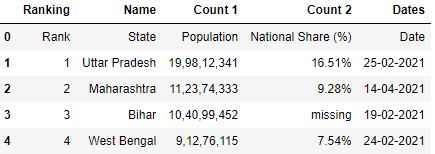
The row 0 seems to be a amend fit for the header. It tin explicate better nigh the figures in the table. You can make this 0 row as a header while reading the CSV past using the header parameter. Header parameter takes the value as a row number.
Annotation: Row numbering starts from 0 including column header
# Read the csv file with header parameter df = pd.read_csv("data1.csv", header=1) df.head() 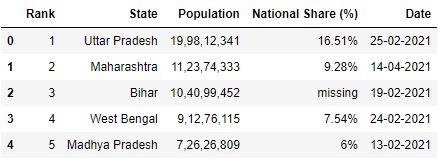
Renaming column headers
While reading the CSV file, you can rename the column headers by using the names parameter. The names parameter takes the list of names of the column header.
# Read the csv file with names parameter df = pd.read_csv( "data.csv" , names=[ 'Ranking' , 'ST Name' , 'Pop' , 'NS' , 'D' ]) df.head() 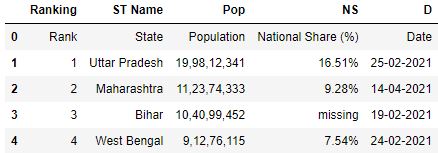
To avoid the former header existence inferred every bit a row for the information frame, y'all tin can provide the header parameter which will override the old header names with new names.
# Read the csv file with header and names parameter df = pd.read_csv( "data.csv" , header=0, names=[ 'Ranking' , 'ST Name' , 'Pop' , 'NS' , 'D' ]) df.head() 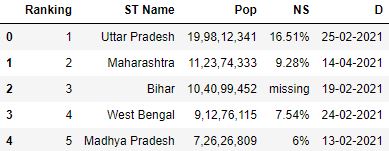
Loading CSV without column headers in pandas
There is a take chances that the CSV file you lot load doesn't accept any column header. The pandas will make the first row as a cavalcade header in the default case.
# Read the csv file df = pd.read_csv("data3.csv") df.head() 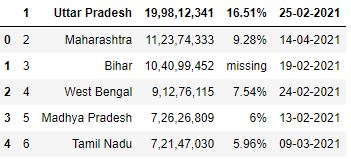
To avoid any row being inferred every bit column header, you can specify header as None. It will force pandas to create numbered columns starting from 0.
# Read the csv file with header=None df = pd.read_csv("data3.csv", header=None) df.head() 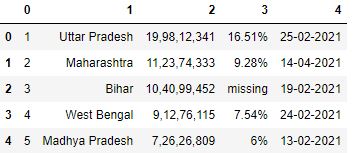
Adding Prefixes to numbered columns
Yous can besides requite prefixes to the numbered column headers using the prefix parameter of pandas read_csv office.
# Read the csv file with header=None and prefix=column_ df = pd.read_csv("data3.csv", header=None, prefix='column_') df.head() 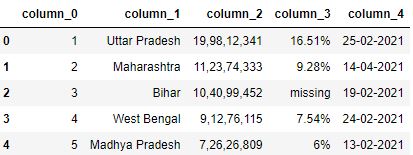
Set any cavalcade(southward) as Index
By default, Pandas adds an initial alphabetize to the data frame loaded from the CSV file. Y'all tin can control this beliefs and make any column of your CSV as an alphabetize by using the index_col parameter.
It takes the name of the desired column which has to be made equally an index.
Instance 1: Making one column equally index
# Read the csv file with 'Rank' as alphabetize df = pd.read_csv("data.csv", index_col='Rank') df.head() 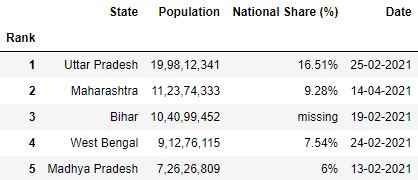
Case 2: Making multiple columns as alphabetize
For ii or more columns to exist fabricated as an index, pass them every bit a list.
# Read the csv file with 'Rank' and 'Date' as index df = pd.read_csv("information.csv", index_col=['Rank', 'Date']) df.head() 
Selecting columns while reading CSV
In do, all the columns of the CSV file are non important. Y'all tin select only the necessary columns later on loading the file but if yous're aware of those beforehand, you lot can save the space and time.
usecols parameter takes the list of columns you desire to load in your data frame.
Selecting columns using list
# Read the csv file with 'Rank', 'Engagement' and 'Population' columns (list) df = pd.read_csv("information.csv", usecols=['Rank', 'Appointment', 'Population']) df.head() 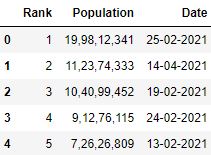
Selecting columns using callable functions
usecols parameter can also take callable functions. The callable functions evaluate on column names to select that specific cavalcade where the part evaluates to True.
# Read the csv file with columns where length of column proper noun > 10 df = pd.read_csv("data.csv", usecols=lambda x: len(x)>10) df.caput() 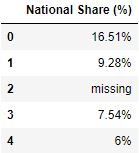
Selecting/skipping rows while reading CSV
You tin skip or select a specific number of rows from the dataset using the pandas.read_csv office. There are three parameters that can practice this task: nrows, skiprows and skipfooter.
All of them have unlike functions. Let's discuss each of them separately.
A. nrows : This parameter allows you to control how many rows you desire to load from the CSV file. It takes an integer specifying row count.
# Read the csv file with 5 rows df = pd.read_csv("data.csv", nrows=v) df 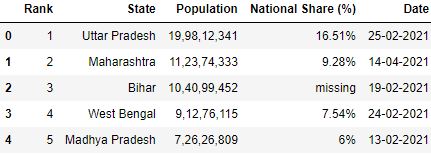
B. skiprows : This parameter allows y'all to skip rows from the showtime of the file.
Skiprows past specifying row indices
# Read the csv file with showtime row skipped df = pd.read_csv("information.csv", skiprows=1) df.caput() 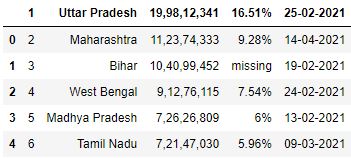
Skiprows by using callback office
skiprows parameter can as well take a callable function as input which evaluates on row indices. This means the callable part will cheque for every row indices to decide if that row should exist skipped or not.
# Read the csv file with odd rows skipped df = pd.read_csv("data.csv", skiprows=lambda x: ten%2!=0) df.head() 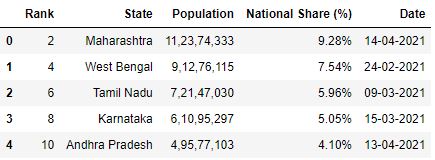
C. skipfooter : This parameter allows you lot to skip rows from the terminate of the file.
# Read the csv file with 1 row skipped from the stop df = pd.read_csv("information.csv", skipfooter=ane) df.tail() 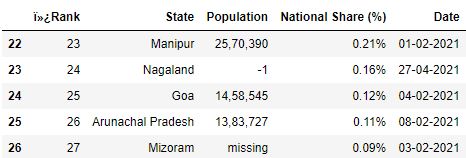
Changing the data type of columns
You lot tin specify the data types of columns while reading the CSV file. dtype parameter takes in the dictionary of columns with their data types defined. To assign the information types, you can import them from the numpy package and mention them against suitable columns.
Information Type of Rank before change
# Read the csv file df = pd.read_csv("data.csv") # Brandish datatype of Rank df.Rank.dtypes dtype ('int64') Data Type of Rank later change
# import numpy import numpy as np # Read the csv file with data type specified for Rank. df = pd.read_csv("data.csv", dtype={'Rank':np.int8}) # Brandish datablazon of rank df.Rank.dtypes dtype ('int8') Parse Dates while reading CSV
Appointment time values are very crucial for information analysis. You can convert a column to a datetime type column while reading the CSV in 2 means:
Method 1. Make the desired column as an alphabetize and pass parse_dates=Truthful
# Read the csv file with 'Engagement' as index and parse_dates=True df = pd.read_csv("data.csv", index_col='Date', parse_dates=True, nrows=five) # Display alphabetize df.alphabetize DatetimeIndex(['2021 -02 -25', '2021 -04 -14', '2021 -02 -19', '2021 -02 -24', '2021 -02 -13'], dtype='datetime64[ns]', name='Date', freq=None) Method 2. Pass desired column in parse_dates as list
# Read the csv file with parse_dates=['Appointment'] df = pd.read_csv("data.csv", parse_dates=['Engagement'], nrows=five) # Display datatypes of columns df.dtypes Rank int64 State object Population object National Share (%) object Date datetime64[ns] dtype: object Adding more than NaN values
Pandas library tin can handle a lot of missing values. But at that place are many cases where the data contains missing values in forms that are not nowadays in the pandas NA values list. Information technology doesn't understand 'missing', 'not found', or 'not available' every bit missing values.
So, y'all demand to assign them equally missing. To do this, use the na_values parameter that takes a list of such values.
Loading CSV without specifying na_values
# Read the csv file df = pd.read_csv("data.csv", nrows=v) df 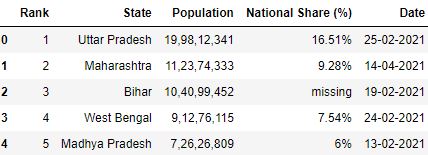
Loading CSV with specifying na_values
# Read the csv file with 'missing' as na_values df = pd.read_csv("data.csv", na_values=['missing'], nrows=5) df 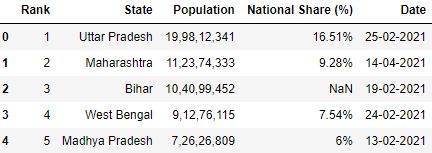
Convert values of the column while reading CSV
You tin can transform, modify, or catechumen the values of the columns of the CSV file while loading the CSV itself. This can be washed by using the converters parameter. converters takes in a dictionary with keys equally the column names and values are the functions to be applied to them.
Let's catechumen the comma seperated values (i.e nineteen,98,12,341) of the Population cavalcade in the dataset to integer value (199812341) while reading the CSV.
# Function which converts comma seperated value to integer toInt = lambda 10: int(10.supplant(',', '')) if x!='missing' else -1 # Read the csv file df = pd.read_csv("information.csv", converters={'Population': toInt}) df.caput() 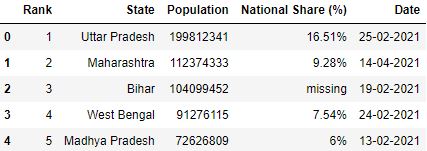
Practical Tips
- Before loading the CSV file into a pandas data frame, always take a skimmed look at the file. It will assist you estimate which columns yous should import and determine what data types your columns should take.
- You should also lookout man for the total row count of the dataset. A system with 4 GB RAM may not exist able to load 7-8M rows.
Examination your knowledge
Q1: Yous cannot load files with the $ separator using the pandas read_csv function. Truthful or False?
Answer:
Answer: False. Because, you lot can use sep parameter in read_csv function.
Q2: What is the employ of the converters parameter in the read_csv function?
Reply:
Reply: converters parameter is used to change the values of the columns while loading the CSV.
Q3: How will you make pandas recognize that a particular cavalcade is datetime type?
Respond:
Respond: By using parse_dates parameter.
Q4: A dataset contains missing values no, non available, and '-100'. How volition you specify them as missing values for Pandas to correctly interpret them? (Assume CSV file name: example1.csv)
Answer:
Answer: By using na_values parameter.
import pandas as pd df = pd.read_csv("example1.csv", na_values=['no', 'not bachelor', '-100']) Q5: How would y'all read a CSV file where,
- The heading of the columns is in the 3rd row (numbered from i).
- The terminal 5 lines of the file take garbage text and should be avoided.
- Only the cavalcade names whose first letter starts with vowels should be included. Presume they are one discussion only.
(CSV file name: example2.csv)
Respond:
Answer:
import pandas as pd colnameWithVowels = lambda x: x.lower()[0] in ['a', 'e', 'i', 'o', 'u'] df = pd.read_csv("example2.csv", usecols=colnameWithVowels, header=two, skipfooter=5) The commodity was contributed by Kaustubh G and Shrivarsheni
Source: https://www.machinelearningplus.com/pandas/pandas-read_csv-completed/
0 Response to "Python Read Last Line of Csv File"
Postar um comentário